Machine Learning approach for gene expression data analysis
 High-throughput microarray technology is an important and revolutionary technique used in genomics and systems biology to analyze the expression of thousands of genes simultaneously. The popular use of this technique has resulted in enormous repositories of microarray data, for example, the Gene Expression Omnibus (GEO), maintained by the National Center for Biotechnology Information (NCBI). However, an effective approach to optimally exploit these datasets in support of specific biological studies is still lacking. Specifically, an improved method is required to integrate data from multiple sources and to select only those datasets that meet an investigator's interest. In addition, to exploit the full power of microarray data, an effective method is required to determine the relationships among genes in the selected datasets and to interpret the biological meanings behind these relationships.
High-throughput microarray technology is an important and revolutionary technique used in genomics and systems biology to analyze the expression of thousands of genes simultaneously. The popular use of this technique has resulted in enormous repositories of microarray data, for example, the Gene Expression Omnibus (GEO), maintained by the National Center for Biotechnology Information (NCBI). However, an effective approach to optimally exploit these datasets in support of specific biological studies is still lacking. Specifically, an improved method is required to integrate data from multiple sources and to select only those datasets that meet an investigator's interest. In addition, to exploit the full power of microarray data, an effective method is required to determine the relationships among genes in the selected datasets and to interpret the biological meanings behind these relationships.
To address these requirements, we have developed a machine learning based approach that includes: (1) An effective meta-analysis method to integrate microarray data from multiple sources; the method exploits information regarding the biological context of interest provided by the biologists. (2) A novel and effective cluster analysis method to identify hidden patterns in selected data representing relationships between genes under the biological conditions of interest. (3) A novel motif finding method that discovers, not only the common transcription factor binding sites of co-regulated genes, but also the miRNA binding sites associated with the biological conditions. (4) A machine learning-based framework for microarray data analysis with a web application to run common analysis tasks on online. Web application, running on Linux server, was implemented using Nginx, PHP and MySql, with jQuery, javascript for the front-end; C++, Java and R for the backend.
AI application in Course Scheduling software
 UEH university, established since 1976 in South of Vietnam, is one of the the top 10 largest universities of the country. Serving more than 40,000 students of undergraduate and graduate programs in multiple majors each year, with 500 faculty members and 200 classrooms located on 7 campuses around the city, the university had a challenging problem of setting up an efficient schedule every semester for effectively using university resources, adapting faculty members' schedules and meeting training programs' requirements. That makes UEH course scheduling problem large and very complicated.
UEH university, established since 1976 in South of Vietnam, is one of the the top 10 largest universities of the country. Serving more than 40,000 students of undergraduate and graduate programs in multiple majors each year, with 500 faculty members and 200 classrooms located on 7 campuses around the city, the university had a challenging problem of setting up an efficient schedule every semester for effectively using university resources, adapting faculty members' schedules and meeting training programs' requirements. That makes UEH course scheduling problem large and very complicated.
In this project, we proposed a new approach for course scheduling problem by combining both AI optimization search methods: Tabu search and genetic algorithm. While genetic algorithm was used to manage problem solutions and to search for optimal solutions, Tabu-search was employed to search for better solutions given a currently achieved or initial one, and to help the genetic algorithm to avoid getting stuck in local optima. We implemented our method in the Facility and Course Scheduling Management system using C++ and MS-Foxpro for Windows. The system had been used at UEH from 2001 to 2006.
Applications of Fuzzy Systems & ANN in time series prediction
 Time series prediction has been widely used in economics, finance, weather forecast, space science, engineering and many other fields due to its benefit in forecasting the future as well as understanding the past. At the Institute of Economics Hochiminh city, it was used for price prediction of some essential merchandises on the market in order to help the City with strategic planning and policy development.
Time series prediction has been widely used in economics, finance, weather forecast, space science, engineering and many other fields due to its benefit in forecasting the future as well as understanding the past. At the Institute of Economics Hochiminh city, it was used for price prediction of some essential merchandises on the market in order to help the City with strategic planning and policy development.
Methods for time series prediction are classified into either local modeling or global modeling type. While the global method utilizes a single set of parameters for the entire forecasting process, local method unitizes different model parameters for different forecasting phases and has been proved performing significantly better than global method. In this project, we combined ANNs with Fuzzy Systems for an effective time series prediction method where ANN took place of global estimator while Fuzzy System was in charge of local estimator. The proposed method was applied to the problem of price prediction of mentioned above.
Machine Learning approach for Business Game software
 Business game software provides virtual real-world environment for students majored in business and management to practice their knowledge, skills and abilities without all the risks that come with a real business. Current state of the art business game software uses mathematic and statistics for modeling of the market, customers, business competitors and business-relevant issues. This approach however has some shortcomings because of the limitation of math and statistics in describing real-world problems, and particularly in constructing the models that can automatically and adaptively learn from historical data.
Business game software provides virtual real-world environment for students majored in business and management to practice their knowledge, skills and abilities without all the risks that come with a real business. Current state of the art business game software uses mathematic and statistics for modeling of the market, customers, business competitors and business-relevant issues. This approach however has some shortcomings because of the limitation of math and statistics in describing real-world problems, and particularly in constructing the models that can automatically and adaptively learn from historical data.
In this project, we proposed an Machine Learning based approach for competitive market modeling using Fuzzy Cognitive Map (FCM), Fuzzy Systems (FZS) and Artificial Neural Networks (ANN). This approach allows a market model, compared with math and statistics based ones, not only to be better in terms of automation and flexibility, but also to provide both market state and market behavior in the simulation environment.
Image feature extraction
to be filled...
HWR Application in academics
to be filled...
OCR Application in academics
to be filled...
An AI heuristic method for Optical Mark Recognition
to be filled...
Social behavior based Online Advertising System on-going project
Traditional Online Advertising Systems use user's contents to determine appropriate ads to deliver.
In this project, we apply machine learning to look for insights from user's contents, particularly those they make available on social networks.
ALU — A connecting platform for UEH Alumni & Students using social media
 Social networks have been around the world for almost two decades, connecting people regardless of location and religion. Use of social networks is an effortless way to join with like-mined people; one is just one click away from an incredible large community of such people. University students use social networks to connect with friends, university staff and family members, sharing their studies, topics of interests, and/or simply their daily activities.
Social networks have been around the world for almost two decades, connecting people regardless of location and religion. Use of social networks is an effortless way to join with like-mined people; one is just one click away from an incredible large community of such people. University students use social networks to connect with friends, university staff and family members, sharing their studies, topics of interests, and/or simply their daily activities.
This project, #CS-2016-02TD sponsored by UEH, creates a platform for connecting university alumni and current students using social media. The alumni and students join the system using their social media account. Using social data analysis of the posts and other information user makes available on the social network, the system discovers the trends and interests of each user and the entire community, set up mentoring sessions and update user with the community upcoming activities, in both professional and entertainment.
Project system was implemented using MySql server for database management, PHP for web programming and C# with Xamarin for mobile app programming. Data analysis was done using Python, R and package libraries, including those for text2vec and gensim. The project website is available online at alu.ueh.edu.vn. The website is currently under a heavy update to comply with new policies from Facebook regarding personal information security.
A pathway based analysis tool for Down syndrome studies
 Human chromosome 21 (HSA21) encodes approximately 160 classical protein coding genes, five microRNAs and an additional >350 genes of unassigned function. Over expression of these genes, as in Down syndrome (DS), will result in complex perturbations of multiple processes involved in neurological development and function. While several recent reports have shown pharmacological rescue of learning and memory deficits in a popular mouse model of DS (the Ts65Dn), this model is trisomic for fewer than 100 HSA21 proteins. Development of safe and effective pharmacotherapies for the cognitive deficits in DS requires an understanding of pathway perturbations in a full trisomy HSA21.
Human chromosome 21 (HSA21) encodes approximately 160 classical protein coding genes, five microRNAs and an additional >350 genes of unassigned function. Over expression of these genes, as in Down syndrome (DS), will result in complex perturbations of multiple processes involved in neurological development and function. While several recent reports have shown pharmacological rescue of learning and memory deficits in a popular mouse model of DS (the Ts65Dn), this model is trisomic for fewer than 100 HSA21 proteins. Development of safe and effective pharmacotherapies for the cognitive deficits in DS requires an understanding of pathway perturbations in a full trisomy HSA21.
Using the web database for Down syndrome studies, here, I developed a systems neuroscience, pathway-based approach, not only to understand non-HSA21 molecular abnormalities observed in DS and mouse models, but also to predict additional abnormalities and potential responses of these systems to drug treatments. The subset of pathways relevant to intellectual disability (ID) in DS are termed DS-ID pathways and are defined as those pathways with components and/or interacting proteins that include HSA21 proteins and one or more ID proteins, proteins known to be involved in ID from mutation analysis in human subjects. The web application is available online at here, using the second option. Please contact me for more detail on the analysis protocol.
 The last two decades have seen incredible growth of the Internet that now connects more than 450 million of computers worldwide, making them more interactive, exciting and powerful, especially in education where used for Computer-Based Test (CBT). Offering an open networking architecture, Internet could make rather seamlessly communicating and sharing resources and data across operating systems and computing platforms. Recent emerging technologies in computer virtualization, cloud computing, Internet browser capabilities, computer network and high-speed wireless connectivity have removed most practical barriers to networking and cross-platform computing anywhere in the world. However, that does not necessarily mean that, CBT would now be possible anywhere in the world, on demand.
The last two decades have seen incredible growth of the Internet that now connects more than 450 million of computers worldwide, making them more interactive, exciting and powerful, especially in education where used for Computer-Based Test (CBT). Offering an open networking architecture, Internet could make rather seamlessly communicating and sharing resources and data across operating systems and computing platforms. Recent emerging technologies in computer virtualization, cloud computing, Internet browser capabilities, computer network and high-speed wireless connectivity have removed most practical barriers to networking and cross-platform computing anywhere in the world. However, that does not necessarily mean that, CBT would now be possible anywhere in the world, on demand.
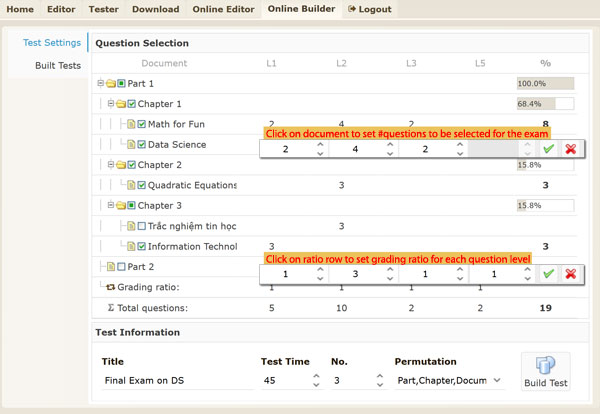
This project aims at designing and developing an Internet based Quiz System using modern technologies to allows CBT online regardless of location and time. The system consists of two main components, one for quiz question and test management, which is used by staff, running either on desktop computer or in cloud; the other for doing test online, which is used by student, running on desktop computer or smart phone as an app, or running in browser as a web app. In addition, a data analysis component will be also integrated to help with analyzing of test results, allowing automatic evaluation of the quiz questions and students' performance.
A web database for Down syndrome studies
 The widely usage of high-throughput microarray technology, an important and revolutionary technique of genomics and systems biology to analyze the expression of thousands of genes simultaneously, has left a problem of optimally exploiting the data from enormous microarray repositories, including the Gene Expression Omnibus (GEO) maintained by the National Center for Biotechnology Information (NCBI) and Connectivity Map (CMAP) maintained by Broad Institute at MIT & Harvard, in support specific biological studies. Specifically, an improved method is required to integrate data from multiple sources and to select only those datasets that meet an investigator's interests.
The widely usage of high-throughput microarray technology, an important and revolutionary technique of genomics and systems biology to analyze the expression of thousands of genes simultaneously, has left a problem of optimally exploiting the data from enormous microarray repositories, including the Gene Expression Omnibus (GEO) maintained by the National Center for Biotechnology Information (NCBI) and Connectivity Map (CMAP) maintained by Broad Institute at MIT & Harvard, in support specific biological studies. Specifically, an improved method is required to integrate data from multiple sources and to select only those datasets that meet an investigator's interests.
This project developed a Human Gene-Protein-Pathway database and system of web interfaces to allow queries against the database online. Data in the database are combined from popular bioinformatics repositories, including NCBI, UniProt and GO for the list of genes, molecules and biological annotations; BIND, HPRD, BioGRID and IntAct for physical protein-protein interactions; KEGG, Reactome, BioCarta and PID for curated pathways of known biological processes. Website development uses PHP for server-side web programming; jQuery and javascript for client-side; and MySql for database management. Client pull technique was also implemented to support large and time consuming queries.
TCP/IP and Windows based network Quiz system
 Development of the TCP/IP protocols widened networking capabilities in the mid of 1990s. TCP/IP offered an open networking architecture that could rather seamlessly communicate and share resources and data across operating systems and computing platforms. Since the late 1990s, introduction of Windows '95, and later Windows NT, had advanced utilization of TCP/IP on network PC, making possible TCP/IP networked CBT to benefit the use of Windows desktops and networks at educational institutes.
Development of the TCP/IP protocols widened networking capabilities in the mid of 1990s. TCP/IP offered an open networking architecture that could rather seamlessly communicate and share resources and data across operating systems and computing platforms. Since the late 1990s, introduction of Windows '95, and later Windows NT, had advanced utilization of TCP/IP on network PC, making possible TCP/IP networked CBT to benefit the use of Windows desktops and networks at educational institutes.
This project designed and developed EmpTest, a Quiz System for CBT on network of computers using TCP/IP protocols, Windows network for example. EmpTest had been used at the University of Economics from 2000 to present. The peak usage of EmpTest is mid of 2000s, more than four times per year and around 5000 students participating each time. When posted to the Vietnam Mistry of Education and Training website for educational software, EmpTest got two millions of downloads in just a month, making it the most popular Quiz software in Vietnam. EmpTest was developed in C++ with Microsoft Visual Studio and Microsoft Foundation Classess (MFC) for Windows applications, using MS-Access for database management. The software website is at emp.tinyray.com. A version of program source code is available online at here.
IPX/SPX Network based Quiz system
 Computer-based testing (CBT) utilizes software programs and technology to deliver assessment test on computer without paper and pencil. It has been widely used by numerous universities and educational institutes for student assessment thanks to its advantages over paper-based testing, both for states that run assessment programs and for students who participate in them. These advantages are recognized by many countries, including the US, encouraged the development of CBT and supporting systems. In addition, with the use of computer network, CBT can be administered on networked PC workstations synchronously, allowing securing the test data and automatically collecting of test results.
Computer-based testing (CBT) utilizes software programs and technology to deliver assessment test on computer without paper and pencil. It has been widely used by numerous universities and educational institutes for student assessment thanks to its advantages over paper-based testing, both for states that run assessment programs and for students who participate in them. These advantages are recognized by many countries, including the US, encouraged the development of CBT and supporting systems. In addition, with the use of computer network, CBT can be administered on networked PC workstations synchronously, allowing securing the test data and automatically collecting of test results.
This project designed and developed a Quiz System for CBT on network of computers using IPX/SPX protocols, Novell network for example. The system had been used at the University of Economics from 1996 to 2000, dozen times per year and around 2000 students participating each time. For system software development, Assembly language was used for IPX/SPX network and low-level GUI programming, and Pascal language for implementation of system interfaces. The program source code is available online at here.